 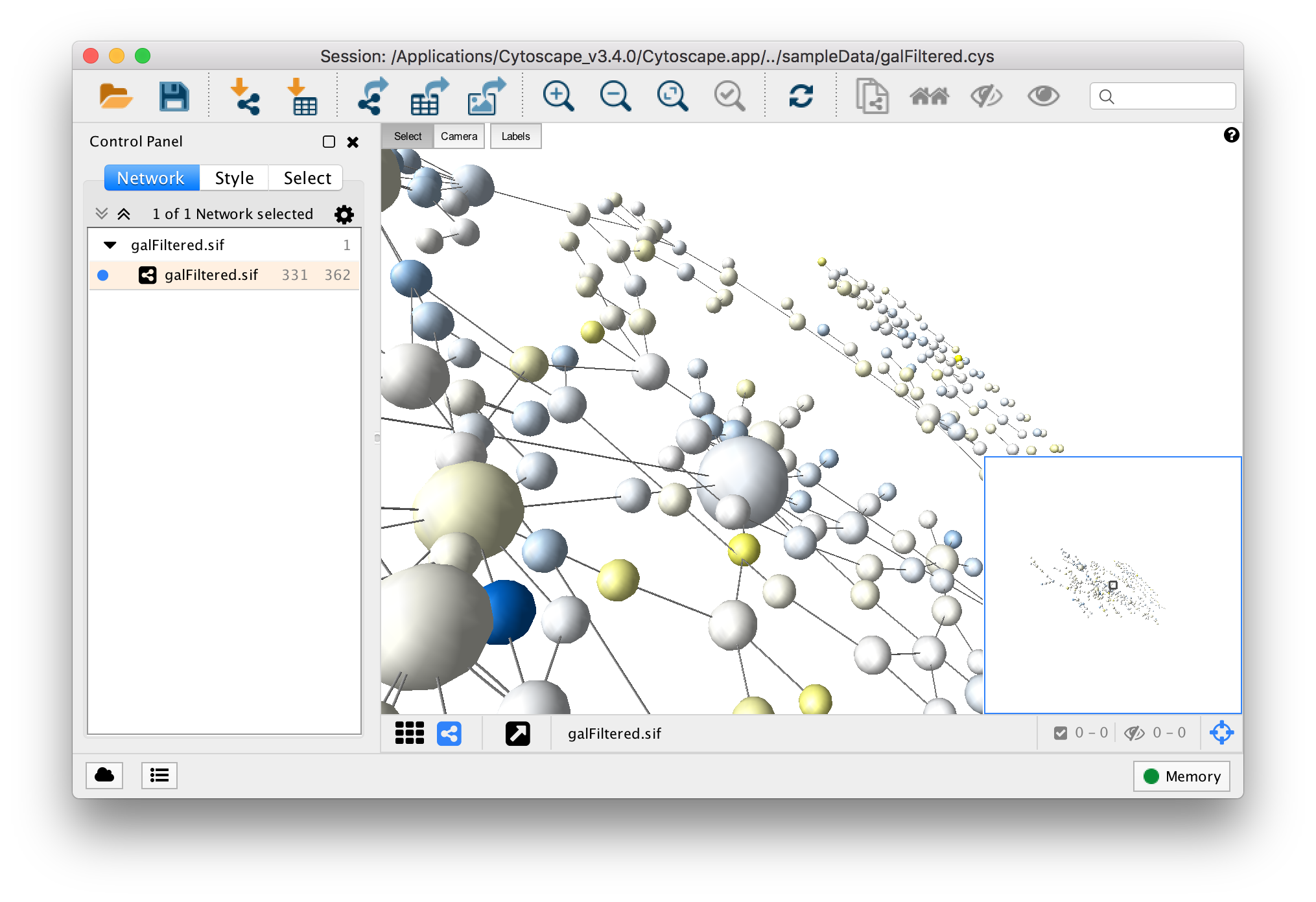
The central organizing metaphor of Cytoscape is a network (graph), with genes, proteins, and molecules represented as nodes and interactions represented as links, i.e. Cytoscape allows the visual integration of the network with expression profiles, phenotypes, and other molecular state information and to link the network to databases of functional annotations. 
Although applicable to any system of molecular components and interactions, Cytoscape is most powerful when used in conjunction with large databases of protein-protein, protein-DNA, and genetic interactions that are increasingly available for humans and model organisms. 
The Core is extensible through a plug-in architecture, allowing rapid development of additional computational analyses and features.Ĭytoscape's roots are in Systems Biology where it is used for integrating biomolecular interaction networks with high-throughput expression data and other molecular state information. A software “Core” provides basic functionality to layout and query the network and to visually integrate the network with state data. Table of Contents Cytoscape 2.4.1 User Manual Introduction Launching Cytoscape (1) If necessary, install Java (2) Install Cytoscape (3) Launch the application Quick Tour of Cytoscape The Menus Network Management The Network Overview Window Command Line Arguments Cytoscape Preferences Managing Properties Managing Bookmarks Managing Proxy Servers Creating Networks Import Fixed-Format Network Files Import Free-Format Table Files Edit a New Network Supported Network File Formats SIF Format GML Format XGMML Format SBML (Systems Biology Markup Language) Format BioPAX (Biological PAthways eXchange) Format PSI-MI Format Delimited Text Table and Excel Workbook Node and Edge Attributes Cytoscape Attribute File Format Import Attribute Table Files Loading Gene Expression (Attribute Matrix) Data Data File Format Example Navigation and Layout Basic Network Navigation Automatic Layout Algorithms Manual Layout Visual Styles Introduction to Visual Styles Visual Attributes, Graph Attributes and Visual Mappers Visual Styles Tutorials Managing Visual Styles Bypassing Visual Styles Finding and Filtering Nodes and Edges QuickFind Filters Select Menu Editing Networks CytoPanels Rendering Engine Annotation Ontology and Annotation File Format Import Ontology and Annotation Custom Annotation Files for Ontologies Other than GO (for Advanced Users) Linkout Acknowledgements Appendix A: Old Annotation Server Format Building your own annotation files Load Data into Cytoscape Getting and Reformatting GO Data Appendix B: GNU Lesser General Public LicenseĬytoscape is a project dedicated to building open-source network visualization and analysis software. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
